- navigate_next Services
- navigate_next About SPILLO-PBSS
- navigate_next Experimental validations / Publications
Invited Research Seminar - School of Medicine and Surgery
TITLE OF THE TALK: A step forward in Drug Development and Personalized Medicine: identification of secondary targets responsible for drug side effects by SPILLO-PBSS
The property of a drug to act with multiple molecular targets usually represents a major drawback both in the drug development process, where toxicity is responsible for the high failure rate (‘attrition rate problem’) of drug candidates, and in medicine, where therapeutic agents often lead to side effects that can cause dose limitation or even discontinuation of treatment.
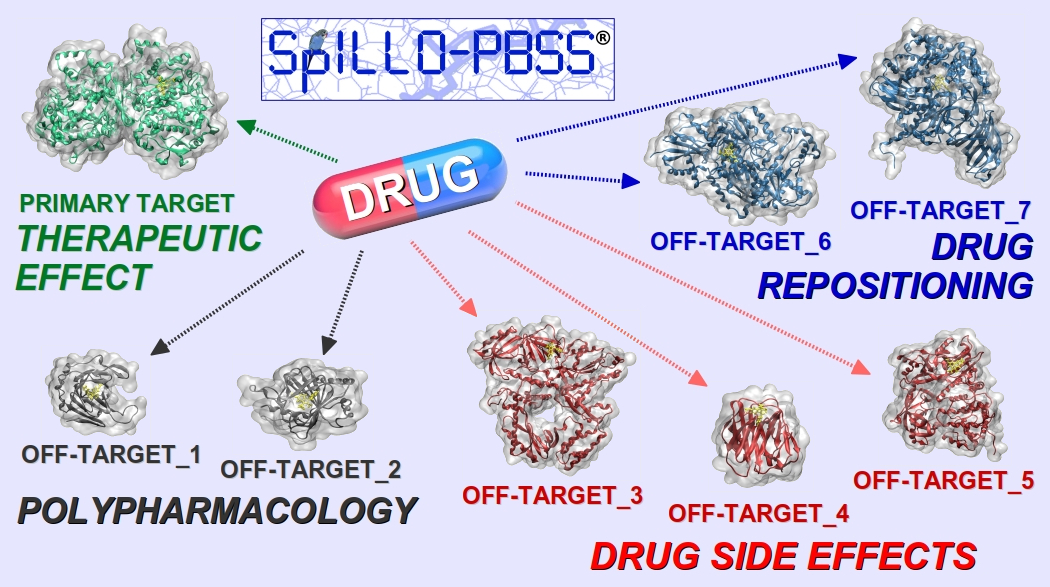
The innovative software SPILLO-Potential Binding Sites Searcher (SPILLO-PBSS)[1] will be presented. It is designed to identify the biomolecular origin of adverse or therapeutic effects of small molecules by a fast identification of their target proteins on a proteome-wide scale (e.g., the whole available human structural proteome). The uniqueness of SPILLO-PBSS lies in its ability to find the potential binding sites on protein 3D-structures even if they are highly distorted and/or apparently inaccessible (e.g., closed, buried, occupied) to the small molecule with respect to an ideal conformation suitable for the binding. In comparison to the other traditional in silico approaches, SPILLO-PBSS is more likely to find proteins responsible for side-effects and/or for additional positive effects of the small molecule (Polypharmacology - Drug repositioning).
The possibility to generate a clearer map of the drug-target interactions has remarkable implications both in the drug development process and in personalized medicine, where treatments with the lowest toxic impact may be selected taking into account individual protein level differences.
REFERENCE:
[1] A. Di Domizio et al (2014). SPILLO-PBSS: Detecting hidden binding sites within protein 3D-structures through a flexible structure-based approach. J. Comp. Chem. 35 (27): 2005-2017; DOI: 10.1002/jcc.23714